The Shyft api
Setup environment for tutorials
Introduction
At its core, Shyft provides functionality through an API (Application Programming Interface). Core functionality of Shyft is available through this API.
We begin the tutorials by introducing the API as it provides the building blocks for the framework. We recommend at least working through the first of these examples. Once you have an understanding of the basic api features, you can move toward configured runs that make use of orchestation. To make use of configured runs, you need to understand how we ‘serialize’ configurations and input data through repositories.
In a separate run_nea_nidelva tutorial, we cover conducting a very simple simulation of an example catchment using configuration files. This is a typical use case, but assumes that you have a model well configured and ready for simulation. In practice, one is interested in working with the model, testing different configurations, and evaluating different data sources. This is in fact a key idea of Shyft – to make it simple to evaluate the impact of the selection of model routine on the performance of the simulation.
In this notebook we walk through a very simple lower level paradigm of using the Shyft ``api`` directly to conduct an individual simulation.
1. Loading required python modules and setting path to Shyft installation
%matplotlib inline
import os
import sys
import datetime as dt
import numpy as np
from matplotlib import pyplot as plt
from netCDF4 import Dataset
# try to auto-configure the path. This will work in the case
# that you have checked out the doc and data repositories
# at same level. Make sure this is done **before** importing shyft
shyft_data_path = os.path.abspath("/workspaces/shyft-data")
if os.path.exists(shyft_data_path) and 'SHYFT_DATA' not in os.environ:
os.environ['SHYFT_DATA']=shyft_data_path
# shyft should be available either by it's install in python
# or by PYTHONPATH set by user prior to starting notebook.
# If you have cloned the repositories according to guidelines:
# shyft_path=os.path.abspath('../../../shyft')
# sys.path.insert(0,shyft_path)
from shyft import hydrology as api
import shyft.hydrology.pt_gs_k
from shyft.hydrology import shyftdata_dir
from shyft.time_series import UtcTimeVector,TimeAxis,TimeSeries,POINT_AVERAGE_VALUE
2. Build a Shyft model
The first point of simulation is to define the model that you will create. In this example, we will use Shyft’s pure api approach to create a model from scratch.
The simulation domain
What is required to set up a simulation? At the most basic level of Shyft, we need to define the simulation domain / geometry. Shyft does not care about the specific shape of the cells. Shyft just needs a ‘geocentroid location’ and an area. We will create a container of this information as a first step to provide to one of Shyft’s model types later.
We are going to be working with the data from the Nea-Nidelva catchment, example dataset. This is available in the shyft-data repository. Above, you should have set your SHYFT_DATA environment variable to point to this directory so that we can easily read the data.
The first thing to do is to take a look at the geography of our cells.
# load the data from the example datasets
cell_data = Dataset( os.path.join(shyftdata_dir, 'netcdf/orchestration-testdata/cell_data.nc'))
# plot the coordinates of the cell data provided
# fetch the x- and y-location of the cells
x = cell_data.variables['x'][:]
y = cell_data.variables['y'][:]
z = cell_data.variables['z'][:]
cid = cell_data.variables['catchment_id'][:]
# and make a quick catchment map...
# using a scatter plot of the cells
fig, ax = plt.subplots(figsize=(15,5))
cm = plt.cm.get_cmap('rainbow')
elv_col = ax.scatter(x, y, c=z, marker='.', s=40, lw=0, cmap=cm)
# cm = plt.cm.get_cmap('gist_gray')
# cid_col = ax.scatter(x, y, c=cid, marker='.', s=40, lw=0, alpha=0.4, cmap=cm)
plt.colorbar(elv_col).set_label('catchment elevation [m]')
# plt.colorbar(cid_col).set_label('catchment indices [id]')
plt.title('Nea Nidelva Catchment')
# print(set(cid))
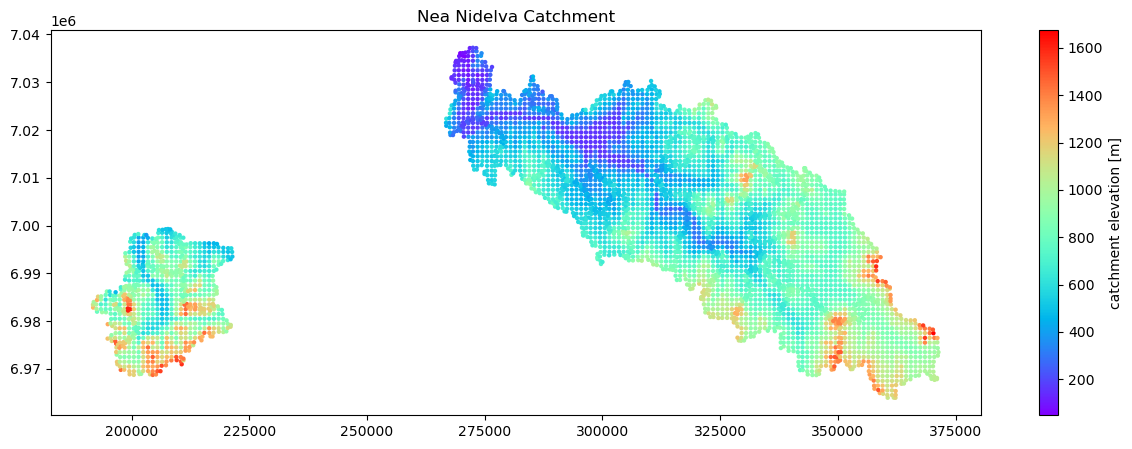
Create a collection of simulation cells
In Shyft we work with ‘cells’, which is the basic simulation unit. In the example netcdf file, we provide the attributes for the cells we are going to plot. But you made need to extract this information from your own GIS, or other data. The essential variables that are minimally required include:
x, generally an easting coordinate in UTM space, [meters]
y, generally a northing coordinate in UTM space, [meters]
z, elevation [meters]
area, the area of the cell, [square meters]
- land cover type fractions (these are float values that sum to 1):
glacier
lake
reservoir
forest
unspecified
catchment_id, an integer to associate the cell with a catchment
a radiation factor (set to 0.9 by default)
If you look at the netcdf file, you’ll see these are included:
print(cell_data.variables.keys())
# output: dict_keys(['x', 'y', 'z', 'crs', 'area', 'forest-fraction', 'reservoir-fraction', 'lake-fraction', 'glacier-fraction', 'catchment_id'])
So the first step is to extract these from the netcdf file, and get them into the model. Note that extracting these values is a pre-processing step that you will likely use a GIS system for. We do not cover that in this notebook.
GeoCellDataVector
In Shyft, there are many custom classes used to optimize the shuffling of data. Frequently you will encounter some type of XxxVector classes. These are essentially lists, and behave as normal python lists. In the following steps we are going to populate this vector with a collection of type api.GeoCellData, which is a custom Shyft type that contains information about the model cells.
Next, we’ll need two other Shyft api classes, the GeoCellData class and the GeoPoint class. These are specialized objects that hold information about each cell. We create these for every cell in the netcdf file. Notice that our netcdf file is quite simple, with one single dimension: cell. So to fill our cell_data_vector, we’ll simply iterate over the cell values in the netcdf file.
# Let's first create a 'container' that will hold all of our model domains cells:
cell_data_vector = api.GeoCellDataVector()
# help(cell_data_vector)
#help(api.GeoPoint)
# from the netcdf file dimensions
num_cells = cell_data.dimensions['cell'].size
for i in range(num_cells):
gp = api.GeoPoint(x[i], y[i], z[i]) # recall, we extracted x,y,z above
cid = cell_data.variables['catchment_id'][i]
cell_area = cell_data.variables['area'][i]
# land fractions:
glac = cell_data.variables['glacier-fraction'][i]
lake = cell_data.variables['lake-fraction'][i]
rsvr = cell_data.variables['reservoir-fraction'][i]
frst = cell_data.variables['forest-fraction'][i]
unsp = 1 - (glac + lake + rsvr + frst)
land_cover_frac = api.LandTypeFractions(float(glac), float(lake), float(rsvr), float(frst), float(unsp))
rad_fx = 0.9
# note, for now we need to make sure we cast some types to pure python, not numpy
geo_cell_data = api.GeoCellData(gp, float(cell_area), int(cid), rad_fx, land_cover_frac)
cell_data_vector.append(geo_cell_data)
# put it all together to initialize a model, we'll use PTGSK
params = api.pt_gs_k.PTGSKParameter()
model = api.pt_gs_k.PTGSKModel(cell_data_vector, params)
The model class
In the case of this tutorial, the model class has been instantiated as a api.pt_gs_k.PTGSKModel. This means that it is using the “PTGSK” model stack. You can instantiate models of type: api.pt_gs_k.PTGSKModel, api.pt_hs_k.PTHSKModel, api.pt_ss_k.PTSSKModel, and api.hbv_stack.HbvModel.
These are central structures in Shyft. All the simulation information is contained within this class.
3. Read forcing data
The next step in the simulation is to get the forcing data. For the tutorials we have provided the data in a series of netcdf files located in the shyft-data repository. Recall that when you set up shyft, you are instructed to clone this repository in parallel to the shyft-doc repository. If you have not done this, then you will have to be sure you have set the SHYFT_DATA environment variable correctly.
Below, we are going to use the shyft.api.ARegionEnvironment class to containerize the forcing data. In this tutorial, this data is station based, but it could be gridded forcing data as well. Once we read the data from the netcdf files, we’ll inject it into the appropriate SourceVectors which are then used for the interpolation step.
re = api.ARegionEnvironment()
# map the variable names in the netcdf file to the source types
source_map = {'precipitation' : ('precipitation.nc', api.PrecipitationSource, re.precipitation),
'global_radiation' : ('radiation.nc', api.RadiationSource, re.radiation),
'temperature' : ('temperature.nc', api.TemperatureSource, re.temperature),
'wind_speed' : ('wind_speed.nc', api.WindSpeedSource, re.wind_speed),
'relative_humidity' : ('relative_humidity.nc', api.RelHumSource, re.rel_hum) }
for var, (file_name, source_type, source_vec) in source_map.items():
nci = Dataset( os.path.join(shyftdata_dir, 'netcdf/orchestration-testdata/' + file_name))
time = UtcTimeVector([int(t) for t in nci.variables['time'][:]])
delta_t = time[1] - time[0] if len(time) > 1 else api.deltahours(1)
time_axis = TimeAxis(time[0], delta_t, len(time))
src_ta= TimeAxis(time,time[-1]+delta_t)
for i in range(nci.dimensions['station'].size):
x = nci.variables['x'][i]
y = nci.variables['y'][i]
z = nci.variables['z'][i]
gp = api.GeoPoint(float(x),float( y), float(z))
data = nci.variables[var][:, i]
dts = TimeSeries(src_ta,data,POINT_AVERAGE_VALUE)
# add it to the variable source vector
source_vec.append(source_type(gp, dts))
nci.close()
# let's take a look at the 'source' data.
# this is exactly the data that will be 'fed' to the interpolation routines
region_environment = re
def plot_station_data(region_environment):
""" plot the data within each source vector of the `ARegionEnvironment`
"""
for fv, sv in region_environment.variables:
n_stn = len(sv)
fig, ax = plt.subplots(figsize=(15,5))
for stn in range(n_stn):
t, d = [dt.datetime.utcfromtimestamp(int(t_.start)) for t_ in sv[stn].ts.time_axis], sv[stn].ts.values
ax.plot(t, d, label=stn)
plt.title(fv)
plt.legend()
plot_station_data(region_environment)
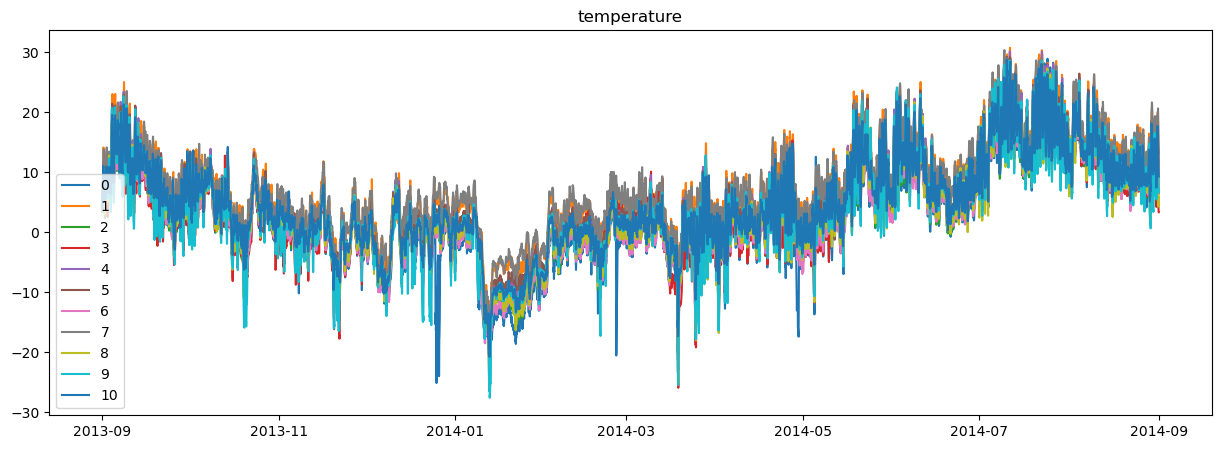

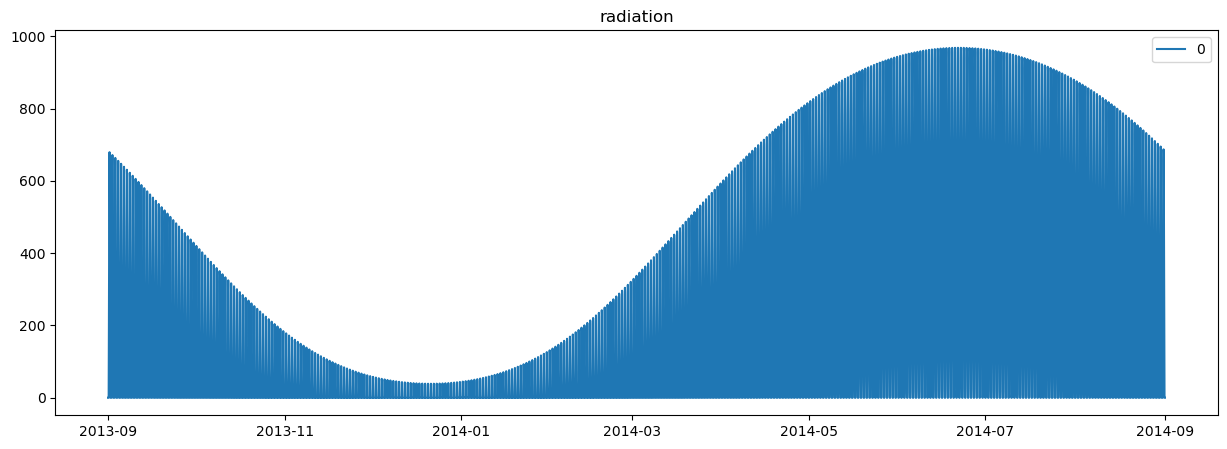
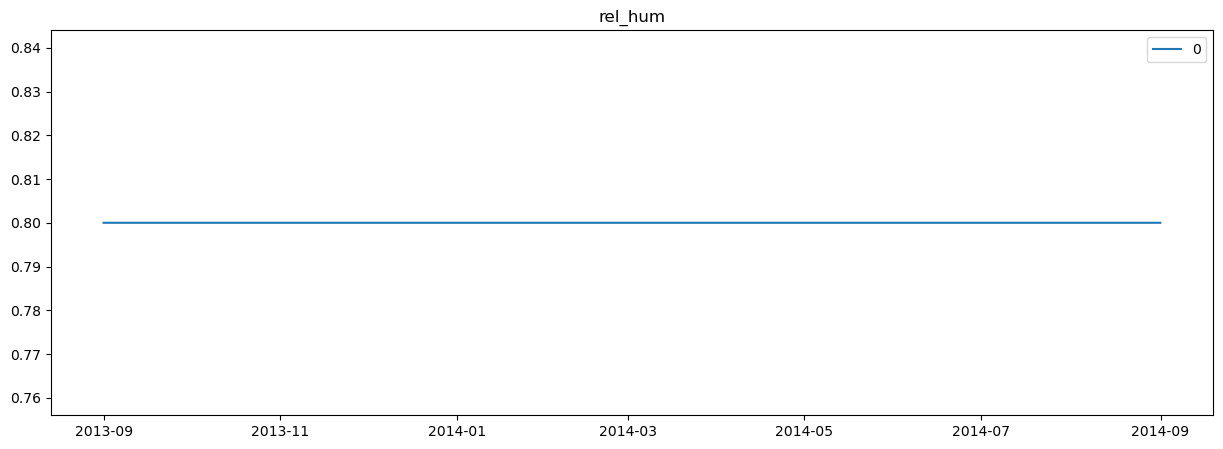
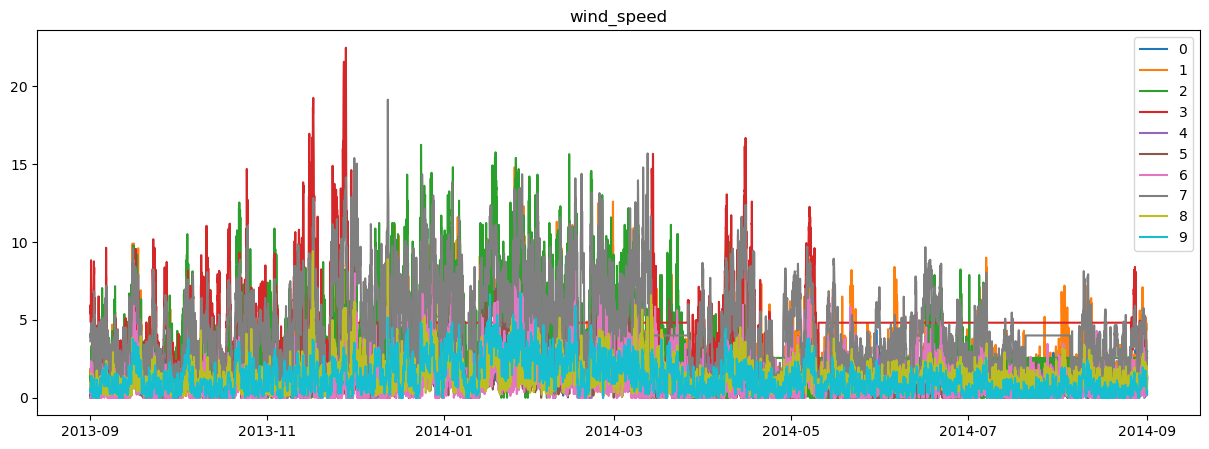
Note that we are simply using a constant relative humidity and simple radiation model. Again, depending on what data you are using, you’ll have different input datasets.
4. Interpolate forcing data to the cells
We are now ready to set up the model interpolation process. In this step, the data is distributed to the cells within your model. Recall that each cell maintains spatial awareness. The interpolation routines within Shyft utilize this information to conduct either Bayesian kriging (temperature) or Inverse Distance Weighting interpolation. See the api documentation, as new methods are being added.
from shyft.time_series import deltahours,Calendar
# next step, distribute the data to the cells
# we need to prepare for the interpolation
# and define the length of the simulation (e.g. time_axis)
# 1. define the time_axis of the simulation:
cal = Calendar()
simulation_ta = TimeAxis(cal.time(2013, 9, 1, 0, 0, 0), deltahours(24), 365)
# 2. interpolate the data from the region_environment to the model cells:
model_interpolation_parameter = api.InterpolationParameter()
model.run_interpolation(model_interpolation_parameter, simulation_ta, region_environment)
# output: True
# at this point we could look at the time series for every cell. Or plot a spatial map...
# TODO: https://data-dive.com/cologne-bike-rentals-interactive-map-bokeh-dynamic-choropleth
from ipywidgets import interact
import numpy as np
from bokeh.io import push_notebook, show, output_notebook
from bokeh.plotting import figure
from bokeh.palettes import viridis
forcing_variables = [fv for fv, sv in region_environment.variables]
output_notebook()
p = figure(title='Forcing Variables', height=300, width=800)
pallette = viridis(100)
cells = model.get_cells()
x = np.array([cell.geo.mid_point().x for cell in cells])
y = np.array([cell.geo.mid_point().y for cell in cells])
def plot_cell_data(model, fv, time_step):
""" plot the data for each cell in the model
Note
-----
This is for demonstration only. Normally one would plot the input data from the
netcdf file directly, but the purpose is to show one how the data is containerized
within Shyft.
"""
if fv == 'temperature': minv, maxv = -40, 40
if fv == 'precipitation': minv, maxv = 0, 400
if fv == 'wind_speed': minv, maxv = 0, 10
else: minv, maxv = -1, 1
def rgb(minv, maxv, val, n, pallette):
dd = np.linspace(minv, maxv, n)
idx = (np.abs(dd - val)).argmin()
return pallette[idx]
# Once we have the cells, we can get their coordinate information
# and fetch the x- and y-location of the cells
data = np.array([getattr(c.env_ts,fv).values[time_step] for c in cells])
colors = [rgb(minv, maxv, d, 100, pallette) for d in data]
r = p.scatter(x, y, radius=300, fill_color=colors, fill_alpha=1, line_color=None)
return p, r
p, r = plot_cell_data(model, 'temperature', 0)
def update(fv='temperature', time_step=0):
plot_cell_data(model, fv, time_step)
push_notebook()
show(p, notebook_handle=True)
interact(update, fv=forcing_variables, time_step=(0,model.time_axis.n))
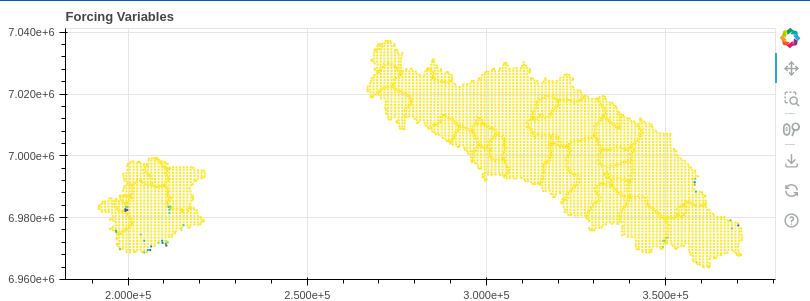
5. Conduct the simulation
We now have a model that is ready for simulation. All the data from our point observations is interpolated to the cells, and we have the env_ts of each cell populated.
The next step is simply to run the simulation.
# #
s0 = api.pt_gs_k.PTGSKStateVector()
for i in range(model.size()):
si = api.pt_gs_k.PTGSKState()
si.kirchner.q = 0.2
s0.append(si)
model.set_states(s0)
model.set_state_collection(-1, True)
As a habit, we have a quick “sanity check” function to see if the model is runnable. Itis recommended to have this function when you create ‘run scripts’.
def runnable(reg_mod):
""" returns True if model is properly configured
**note** this is specific depending on your model's input data requirements """
return all((reg_mod.initial_state.size() > 0, reg_mod.time_axis.size() > 0,
all([len(getattr(reg_mod.region_env, attr)) > 0 for attr in
("temperature", "wind_speed", "precipitation", "rel_hum", "radiation")])))
# run the model, e.g. as you may configure it in a script:
%time
if True: #runnable(model):
model.revert_to_initial_state()
model.run_cells()
else:
print('Something wrong with model configuration.')
# output:
# CPU times: user 2 µs, sys: 0 ns, total: 2 µs
# Wall time: 7.63 µs
Okay, so the simulation was run. Now we may be interested in looking at some of the output. We’ll take a brief summary glance in the next section, and save a deeper dive into the simulation results for another notebook.
6. Take a look at simulation results
The first step will be simply to look at the discharge results for each subcatchment within our simulation domain.
# We can make a quick plot of the data of each sub-catchment
fig, ax = plt.subplots(figsize=(20,15))
# plot each catchment discharge in the catchment_ids
for i,cid in enumerate(model.catchment_ids):
# a ts.time_axis can be enumerated to it's UtcPeriod,
# that will have a .start and .end of type utctimestamp
# to use matplotlib support for datetime-axis, we convert it to datetime (as above)
ts_timestamps = [dt.datetime.utcfromtimestamp(int(p.start)) for p in model.time_axis]
data = model.statistics.discharge([int(cid)]).values
ax.plot(ts_timestamps,data, label = "{}".format(model.catchment_ids[i]))
fig.autofmt_xdate()
ax.legend(title="Catch. ID")
ax.set_ylabel("discharge [m3 s-1]")
# output: Text(0, 0.5, 'discharge [m3 s-1]')
